Life sciences research is in the midst of an extraordinary fruitful period. New technologies are enabling researchers to probe biological phenomena with ever increasing time and spatial resolution, ever greater selectivity, and ever greater coverage. Researchers can sequence thousands of mRNAs and determine their expression levels in single cells, image the development of an embryo in real time, or measure blood flow in the brains of humans performing sophisticated cognitive tasks. These and many other experiments rely on technologies involving complex experimental protocols, sophisticated machinery, and multi-step data analysis pipelines. I believe that this complexity should make us pause and ask: How do we determine whether a technology has reached maturity? How do we determine whether we understand the steps in the experimental protocol well enough?
Consider the case of functional magnetic resonant imaging (fMRI). Magnetic resonant imaging has been routinely used for brain imaging since the early 1990s. Considering the time elapsed, one would expect the fMRI pipeline to be a mature technology. However, just a year ago, Eklund et al. reported that the use of “simplistic temporal autocorrelation model[s]” has resulted in false positive rates of “up to 70%” in fMRI studies.
While this problem has been minimized by some as merely an example of the incompetent use of statistics, I believe that this is a structural – thus, far more serious – problem. Specifically, Eklund et al. demonstrate that the three most popular software packages for fMRI analysis can yield very high false positive rates. This reveals a serious flaw in the multi-step data analysis pipeline commonly used by researchers, and leads me to conclude that fMRI is not yet a mature technology. This is particularly worrisome since fMRI evidence is now being used in courts.
Consider also the case of microarrays, a twenty year old technique to assess DNA sequence and gene expression.3 A search of the Web of Knowledge database reveals that already by the year 2000 more than a thousand original research papers involving microarrays were being published every year (Fig. 1a). However, in the early 2000s, microarrays were plagued by poor quality control in manufacturing and ‘batch effects.’ This would not be too serious a problem if most work on microarrays focused on validation and standardization of the technology. However, from the very start, microarrays were being used for studying diseases and developing treatments.
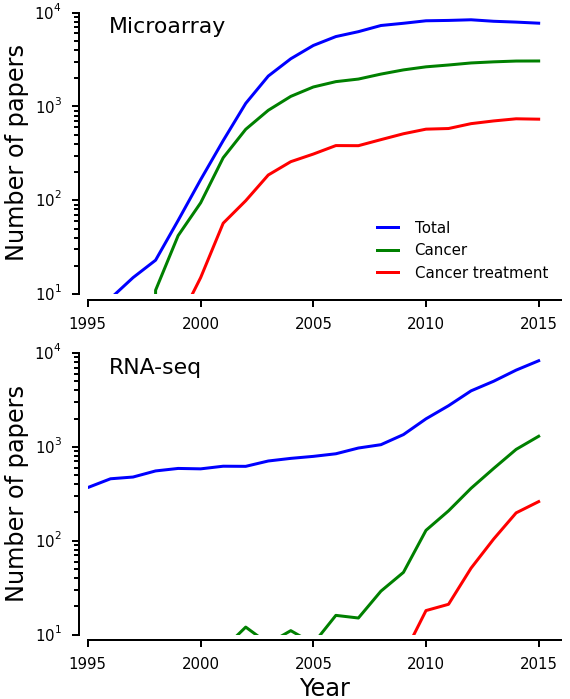 |
Figure 1: Number of papers recorded in ThomsonReuters’ Web of Knowledge involving microarrays (top) or next generation sequencing (bottom) using the terms indicated in the legend. Data was obtained by querying Web of Knowledge for publications filtered by topic and restricting to ‘articles’. Queries performed were: ‘microarray’, ‘microarray AND cancer’, microarray AND cancer AND treatment’, ‘‘next generation sequencing’ OR ‘deep sequencing’ OR ‘RNA-seq‘’, ‘‘next generation sequencing’ OR ‘deep sequencing’ OR ‘RNA-seq’ AND cancer’, and ‘‘next generation sequencing’ OR ‘deep sequencing’ OR ‘RNA-seq’ AND cancer AND treatment’. | ||||||||||||||||||||||||||||||||||||
| - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
In The (Honest) Truth About Dishonesty (page 188), Dan Ariely writes that ‘every time business breaks through new technological frontiers […] such progress allows people to approach the boundaries of both technology and dishonesty.’ In other words, when technologies are in their infancy, (i) innovators pursue many distinct, poorly-validated, directions, (ii) there is overwhelming uncertainty about what can be accomplished with the technology, and (iii) dishonest behavior flourishes. The ‘boundaries of both technology and dishonesty’ were indeed approached with microarrays. First, many labs were pursuing their own ‘home brewed’ approaches that, without verification, were presented as superior to all other approaches. Unfortunately, many labs were actually using inappropriate tools for their data analyses. The most egregious example was perhaps the use of Microsoft Excel without turning off auto-formatting, which led to gene names being turned into dates and lost from the analysis. While potentially due to journal requirements for authors, a recent study estimated that 20% of papers published between 2005 and 2015 still uploaded Excel files with this error.
Second, many labs made extraordinary claims that were later shown to be unfounded. For example, hundreds of studies claimed to have identified multi-gene signatures for different cancers. However, an independent review of 16 studies of multiple gene signatures in non-small cell lung cancer stated “…we found little evidence that any of the reported gene expression signatures are ready for clinical application. We also found serious problems in the design and analysis of many of the studies.”
Third, several players tried to profit from the high expectations. Several companies rushed to market gene expression tests for cancer detection. In 2011, two such companies, LabCorp and Correlogic Systems, were warned by the FDA that they had failed to validate their tests for early detection of ovarian cancer. Unsurprisingly, those companies claimed the tests were “home brews,” and so did not require validation. Showing that academics are not immune to these tendencies, Duke University closed three cancer trials based on fraudulent research by Anil Potti that had been published earlier in Nature Medicine (see NYT article for details).
Costs of adoption of immature technologies
Immature technologies are unreliable tools. During the initial phases of development one should be making experiments on the technology, not using the technology in experiments. Indeed, a scientist can only know that she got the right outcome out of an experiment if she a priori knows that the outcome is correct, or if she knows that the experiment must work because it is a well understood technology. At the scientific frontier, either because of studying new phenomena or because of using an immature technologies, she cannot possible know whether the experimental results are trustworthy or not.
Every scientist is taught that in the long-term science self-corrects. Unfortunately, there is no guarantee that the ‘long-term’ won’t be decades away or that the cost to the scientific enterprise won’t be quite significant (see Ioannidis, Open Consortium and Nissen et al.). One cost is the littering of the scientific literature with irreproducible, un-replicable studies. Lack of standardization and lack of understanding of the steps in a pipeline lead to a lack of appropriate documentation and make it impossible to reproduce results. Leek and Jager estimated that between 20% and 50% of studies in bioinformatics may be irreproducible, whereas Nekrutenko and Taylor estimated that about 86% of studies using high-throughput technologies may be irreproducible. The situation for replicability is at best similar. Mobley et al. surveyed researchers at MD Anderson Cancer Center and found that 55% of respondents had tried and failed to replicate published results, and that 60% could not get necessary information from authors of published studies.
Another significant cost of immature technologies is monetary. Immature technologies are cutting edge and expensive. Muir et al. compiled data from NIH external awards and estimated that in period 2003–2012, over $1 billion a year was spent supporting research using microarrays. Because of concerns about its performance, microarrays have been recently replaced by next generation sequencing (NGS) technologies (Fig. 1b). Since 2008, funding for studies incorporating sequencing technologies has increased dramatically, exceeding $2.5 billion a year by 2014 (Fig 1a of Muir et al.). However, technologies using NGS for estimation of gene expression, such as RNA-seq, are not mature either.
A rapidly growing number of studies are demonstrating how sample preparation, analysis pipelines, and other experimental protocols biased what genetic variation was found, or missed (see Hansen et al. and Alberti et al.), that changing the number of biological replicates or the statistical analysis tools can yield radically different outcomes, and that a critical step in many RNA-seq analyses – extraction of ribosomal RNAs – leads to dramatic changes in estimated counts for many genes, a bias mechanism that is not accounted by current recommended procedures (see Roberts et al. and Adiconis et al.).
Causes of adoption of immature technologies
Considering the risks and high costs of immature technologies, one may wonder why they are being broadly adopted so promptly. I see two main reasons for this. The first and foremost reason, is the wish by every scientist to make exciting new discoveries that will survive into posterity. It is understandable that one would believe that using a new technology will provide an advantage in this quest. Perversely, the fact that immature technologies tend to have higher rates of false positives because of improper use will lead to more ‘extraordinary’ results, which in turn will have a greater chance of being published in a high-profile journal, and therefore generate even more interest in the technology.
The second reason is the requirement by review panels for federal funding program that applicants use the most cutting-edge approaches even when such approaches may not be called for. I have personally observed this as a reviewer and have experienced it as an applicant. Indeed, research suggests that researchers, reviewers, program officers, and journal editors place too high a value on novelty and potential significance and too low a value on the evaluation of limitations and caveats (see Nissen et al. and Ioannidis et al.). Take Nature Methods; it describes itself as “a forum for the publication of novel methods and significant improvements to tried-and-tested basic research techniques in the life sciences”. However, it also places “a strong emphasis on the immediate practical relevance of the work presented.”
It is naïve to expect that complex technologies emerge ready for practical use. History clearly demonstrates that such an expectation is not realistic. Fortunately, some stakeholders are starting to pay attention to this matter. For example, the National Institute of General Medical Sciences (NIGMS) has started two initiatives for supporting technology development. As their announcement states: “Historically, support for technology development has generally been coupled to using the technology to answer a biomedical research question. Although in the later stages of technology development this coupling is often useful, in the early stages it can hinder exploration of innovative ideas that could ultimately have a big impact on research” (see NIGMS Loop ). And journals are also now frequently demanding deposition of code and data used to generate published results.
A way forward
If publishers and research funders truly want to set reproducible research as a goal, then they must make choices that align career incentives with the publication of reproducible research. As NIGMS is now proposing to do, stakeholders must reward the development, validation, and standardization of new tools. In particular, they must recognize the scientific importance and the intellectual merit of not just tool development, but also tool validation, and standardization.
Editors, federal funding agencies, and foundations have more power over what is recognized by the scientific community than they typically acknowledge. If scientific prizes are awarded not only for ‘discoveries’ but also for development of new technologies, then scientist will more readily acknowledge careful development of tools. Indeed, while prizes have historically been mostly awarded for discoveries, technologies – from the microscope to PCR – have had a much greater impact on the growth of our scientific knowledge than most discoveries. This impact is visible in the fact that papers reporting truly significant methods or resources tend to be the most highly-cited.
I believe, however, that the changes under way are not enough. Immature technologies must not be used, for example, for developing assays for early detection of cancer. The perverse incentives of using immature technologies discussed above will always tempt some to use them for short term gain. I see two possible solutions to this conundrum. The first is top-down and has its roots in historical examples, whereas the second is bottom-up and inspired by the organic emergence of the Open Source movement.
In order to motivate the top-down historically-inspired approach, let me recall some facts concerning the development of thermometers. The first thermometers were made in the late 1600s and suffered from low replicability in their construction and the use of non-standard temperature scales. In order to agree on a standard scale, scientists had to agree on the definition of the fixed points for the scale. Those were agreed to be the temperature at which water freezes at standard pressure, and the temperature at which water boils, also at standard pressure. Unfortunately, while trying to rigorously determine the temperature at which water boils, it became apparent that factors such as the amount of air dissolved in the water, the material used for the container, or the degree of cleanliness of the container all strongly altered the boiling temperature. In order to address this matter, The Royal Society tasked a team of scientists, led by Henry Cavendish, to study the matter and recommend guidelines for the setting of the fixed points. Their recommendations were published in the Philosophical Transactions in 1777 and provided an advance that set in motion the development of thermodynamics and the steam engine. Similarly, federal funding agencies could request that The National Research Council assemble teams tasked with the analysis of promising new technologies and the creation of recommendations regarding (i) their suitability for use in research, and (ii) guidelines for their use.
An alternative, bottom-up model for evaluation of new technologies would replicate the approach followed by the Open Source movement. Producing a successful software package involves many aspects: writing code, testing and validating code, writing documentation, developing examples and tutorials, and so on. Many software packages are started by a single person, but successful packages can amass very large development teams. The Linux kernel is perhaps the paradigmatic example. The first version was created solely by Linus Torvald, but the most recent versions include contributions from several thousand developers. The contributions of all those developers are recorded in documents that accompany the distribution of the software and are publicly available.
Importantly, an open source software may have several concurrent distributions available to the community. The different distributions will have different levels of stability (and thus require different levels of sophistication on the part of the end user), and will interact appropriately with different distributions of other software packages. In this way, the technology signals to users where it is on its development curve. It is illuminating to contrast this practice with the situation described by Sinha et al. for the incorporation of a new sample amplification technology into the Illumina HiSeq 4000 sequencing machines.
The cost to the scientific enterprise and to society of publishing research predicated on the use of immature technologies is significant. Action needs to be taken in order to safeguard the mission and the standing of science among the public, especially in a time of alternative facts and anti-science postures. If we do not take action, the scientific enterprise will waste billions of dollars conducting irreproducible research that does not advance scientific knowledge and does not help to improve quality of life in our societies (see Bowen and Casadevall).